PDF) Suspension TRAPping Filter (sTRAP) Sample Preparation for Quantitative Proteomics in the Low µg Input Range Using a Plasmid DNA Micro-Spin Column: Analysis of the Hippocampus from the 5xFAD Alzheimer's Disease Mouse

Comparative proteomics analysis of in-gel digestion and sTRAP protocol... | Download Scientific Diagram
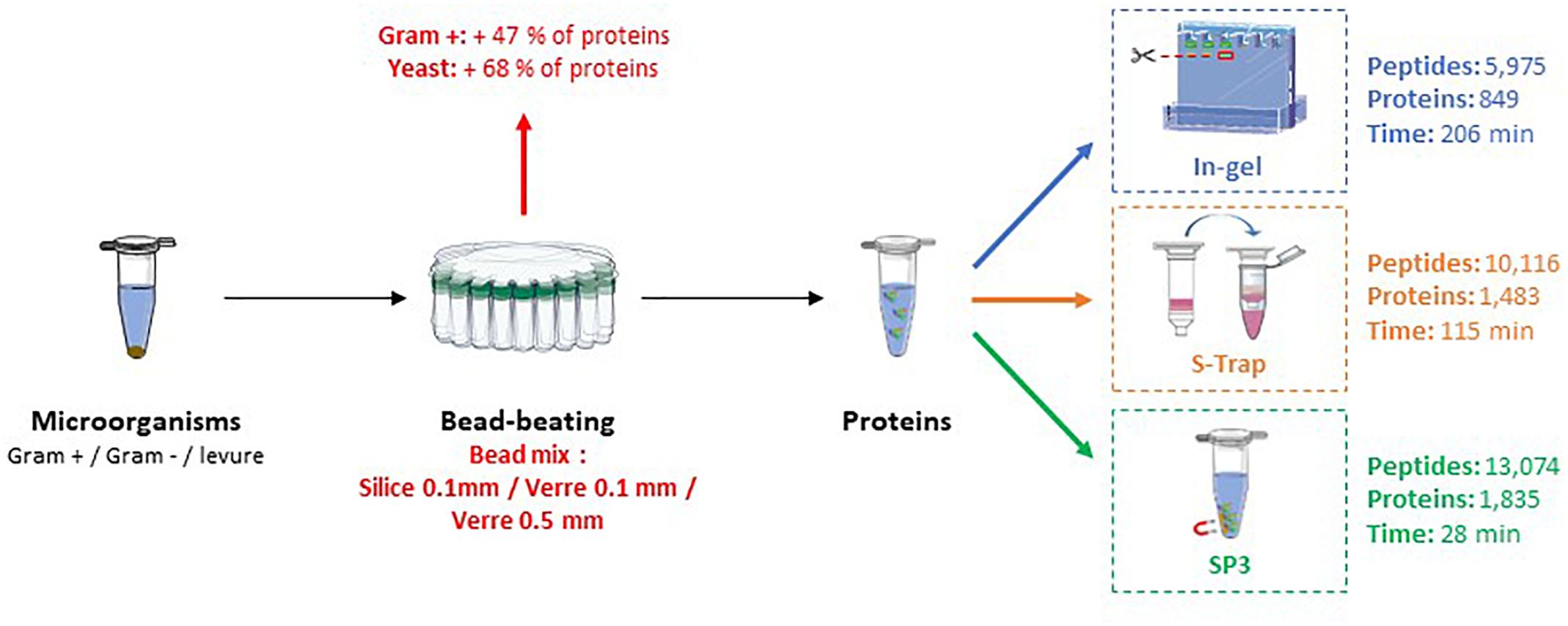
Frontiers | Evaluation of Sample Preparation Methods for Fast Proteotyping of Microorganisms by Tandem Mass Spectrometry

Optimized Suspension Trapping Method for Phosphoproteomics Sample Preparation | Analytical Chemistry
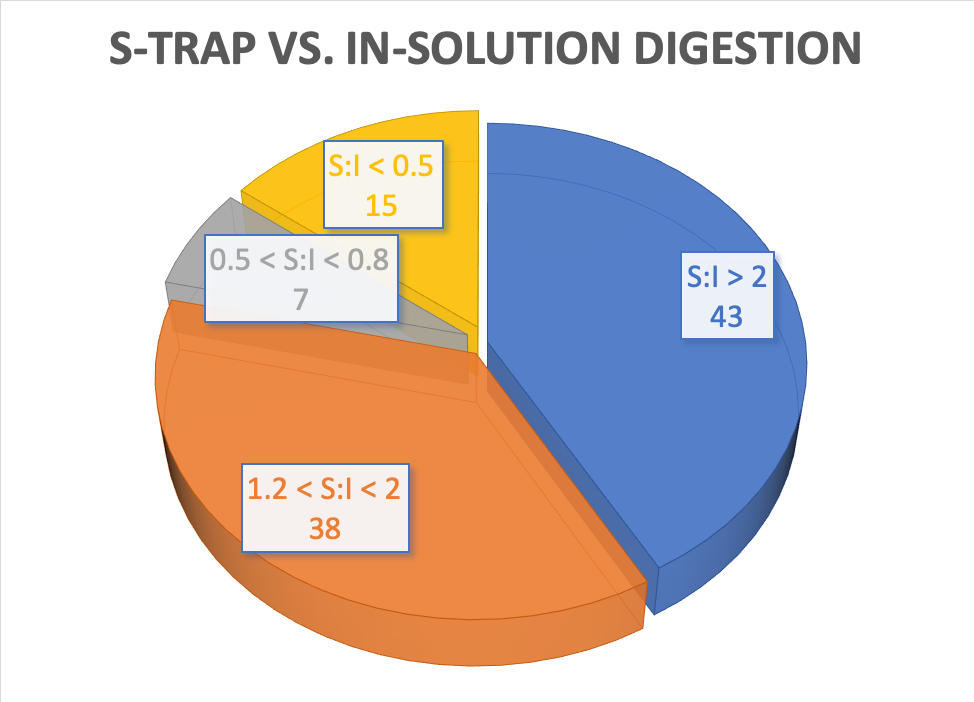
Application of S-Trap in Mass Spectrometry-Based Serum Biomarker Discovery – ISB High School Interns 2019

Suspension Trapping (S-Trap) Is Compatible with Typical Protein Extraction Buffers and Detergents for Bottom-Up Proteomics | Journal of Proteome Research

.png?width=1174&name=Fig%201%20Protifi%20S-Trap%20protocol%20(1).png)